OpenAccess.ai: How Cheap Can We Make Academic Publishing?
What Problem Are We Trying to Solve?
Article Processing Charges at major journals run to thousands of dollars per paper. The editorial infrastructure - submission systems, copyediting, typesetting, hosting - is expensive. What I want to know is how much of the process can be automated, particularly for machine generated academic content.
If you can automate a substantial fraction of the editorial process, (as Preprints.ai is attempting to do) what happens to the unit economics? OpenAccess.ai is an experiment designed to provide some data.
There's a second motivation. OpenScience.ai (in theory) generates research articles. Those articles need somewhere to go. Conventional open access infrastructure is designed for human-authored papers, with submission systems and APC structures that don't map cleanly onto a machine-generated pipeline. OpenAccess.ai is built to accept submissions via API, which means it can serve as the publishing endpoint for the entire experimental pipeline I’m building, without friction.
The Technical Architecture
Core Stack
OpenAccess.ai is a Next.js 14 application deployed on Netlify with Supabase for PostgreSQL, auth, and file storage, and Railway for microservices. Serverless-first, no dedicated servers, minimal ops overhead. If we're testing whether publishing can be radically cheap, the infrastructure costs need to be radically cheap too.
The Document Processing Pipeline
When someone uploads a PDF, the platform needs to extract the full text, identify sections, parse references, pull out figures, and generate structured metadata.
Stage 1: Fast extraction
The upload route runs pdf-parse for basic text extraction in parallel with a Supabase Storage upload. For DOCX files, mammoth.js handles the conversion. This produces something immediately. However, for PDFs with embedded fonts (which is most academic PDFs), pdf-parse returns garbled text.
Stage 2: AI vision extraction
When pdf-parse fails (detected by our garbled-text heuristic that checks for XMP metadata patterns, font encoding artifacts, and low information density), the client triggers a Supabase Edge Function that sends the PDF to Claude's document vision API. Claude Haiku reads the visual layout of each page and produces clean Markdown with proper sections, author lists, figure captions, and references.
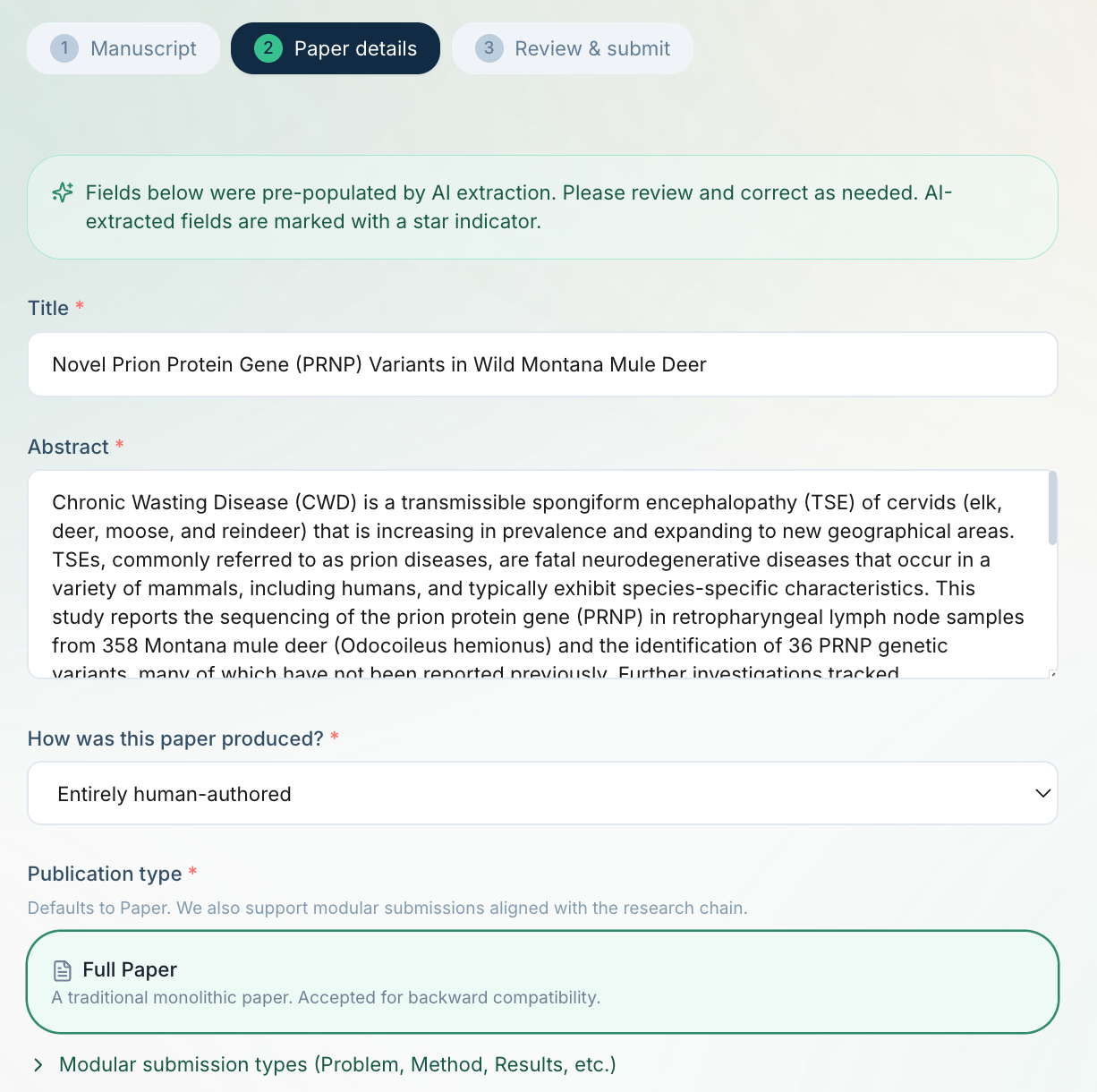
Stage 3: Structured metadata extraction
The extracted text is sent to Claude Haiku again, this time with a structured prompt that returns JSON with title, abstract, authors with affiliations, keywords, and subject area (mapped to bioRxiv's taxonomy of 25 subject areas). This populates the submission form automatically.
Stage 4: Figure extraction
A PyMuPDF-based microservice on Railway extracts embedded images from the PDF, filters by minimum dimensions, and stores them in Supabase Storage. The figure URLs are matched to the document AST by order.
The AST Document Model
All documents are converted to an internal Abstract Syntax Tree inspired by MyST Markdown (itself derived from mdast and Stencila). The AST is the canonical representation - HTML, JATS XML, PDF, and all citation formats are rendered from it on demand.
RootNode
├── meta: { title, abstract, sections[], wordCount, figureCount, referenceCount }
├── children: SectionNode[]
│ ├── SectionNode (id: "introduction")
│ │ ├── HeadingNode
│ │ ├── ParagraphNode[]
│ │ └── FigureNode { url, caption, label }
│ ├── SectionNode (id: "methods")
│ └── SectionNode (id: "results")
└── references: ReferenceNode[] (CSL-JSON format)
Sections are first-class nodes, not just CSS classes on headings. This means you can programmatically extract "the methods section" or "all figure captions" without parsing HTML — which is essential for the AI review pipeline.
Peer Review via Preprints.ai
Every submission is automatically sent to Preprints.ai for assessment. The review system uses 8 AI reviewers that evaluate the paper across multiple dimensions:
- Integrity scoring (A–E grade scale)
- Novelty assessment (1–5 tiers: minimal to groundbreaking)
- Methodology checks (statistical validity, sample sizes, controls)
- Transparency audit (ethics statement, data/code availability, funding disclosure, COI, pre-registration)
- Provenance analysis (for AI-generated papers: model plausibility, prompt injection detection, reproducibility signals)
The grade, strengths, concerns (with severity: critical/major/minor), and per-reviewer scores are stored as structured JSON and displayed on the article page. The review is fully transparent — readers see exactly what the AI reviewers found, not just a binary accept/reject.
Output Formats and Academic Standards
A publishing platform is only useful if it produces outputs that the academic ecosystem can consume. OpenAccess.ai generates:
For Indexing and Discovery
- JATS XML - Journal Article Tag Suite, the standard for CrossRef DOI deposit, PubMed indexing, and PMC archiving. Generated on-demand from the AST via astToJats().
- Dublin Core meta tags - for library catalog discovery (Zotero, Mendeley)
- Highwire Press tags - citation_title, citation_author, etc. for Google Scholar
- Schema.org JSON-LD - ScholarlyArticle type with full provenance metadata
For Citation
- Citation styles: APA 7th, Vancouver, Chicago, Harvard, MLA 9th, BibTeX, RIS - generated client-side from article metadata using Citation Style Language conventions
- CFF (Citation File Format) - for software-like citations
For Machine Learning and Linked Data
- Croissant ML - ML dataset metadata standard, making the corpus discoverable for training pipelines
- RO-Crate - Research Object packaging with full provenance
- W3C DCAT - Data Catalog Vocabulary for dataset registries
- DataCite JSON - DOI registration metadata
Persistent Identifiers
- Handle System - Each article gets a globally unique persistent identifier (e.g., 20.500.14906/oai.biology.2026.00001) registered via Handle.net's REST API
The Article Reader
The article page is modelled on eLife's reading experience
- Tabbed interface. Full text, Figures & data, Peer review (with full Preprints.ai assessment), Summary
- Sticky section navigation. Left sidebar with table of contents generated from AST section nodes, highlights current section on scroll
- Inline citations. Hover to see reference details, click to scroll to bibliography
- AI-powered Q&A. Claude-powered chat that answers questions about the article content
- Branded PDF export: Print-optimized HTML with OpenAccess.ai header, proper title block, A4 pagination
- Article metrics. Views, downloads, citations tracked per article
Authentication
Users sign in via ORCID OAuth or GitHub OAuth. ORCID profiles are enriched with works count and affiliation data. The affiliation field uses the ROR API (Research Organization Registry) for standardised institution lookup.
Agent-First Architecture
OpenAccess.ai exposes an MCP (Model Context Protocol) endpoint at /api/mcp with tools for:
- submit_paper - Programmatic manuscript submission (API key + credits)
- check_submission - Query review status
- get_article - Fetch in any format (JSON, JSON-LD, BibTeX, Croissant, etc.)
- search_articles - Full-text search
- get_subject_areas - Valid taxonomy lookup
Plus a REST API documented at /api/openapi.json and discoverable via /llms.txt, .well-known/ai-plugin.json, and an agent card. The API uses credit-based billing: generate an API key, purchase credits via Stripe, spend 1 credit per submission.
This is what enables OpenScience.ai to generate a paper and submit it to OpenAccess.ai without any human in the loop.
Tools That Inspired the Platform
Several open-source projects shaped the architecture:
What We Can Look to Do Next
We need to run a meaningful volume of articles through the full pipeline - generate via OpenScience.ai, review via Preprints.ai, publish via OpenAccess.ai - and measure the per-article cost at each stage.
Figure extraction needs improvement. The PyMuPDF service extracts embedded images but doesn't yet match them to figure captions reliably. We're evaluating eLife's enhanced-preprints-image-server (IIIF-based) for production-grade image delivery.
Reference enrichment is the next major quality improvement. Replacing AnyStyle with jats-ref-refinery would dramatically improve citation linking by querying four external databases rather than relying on a single CRF parser.
Indexing and registration remain open questions. A journal that publishes AI-generated research will not easily get into PubMed or Dimensions.ai. But understanding what metadata, what provenance signals, and what quality documentation would be required for AI-generated research to be treated as a first-class research output is itself a useful question for the field to be working on.
The most interesting longer-term question is whether a hybrid editorial model makes sense - AI review as the default pathway, with human editorial escalation for papers that score ambiguously or that make claims in domains where agent reviewers have known weaknesses.If you're working on any of these problems and want to compare notes, or if you spot something obviously wrong with the approach, I'd be glad to hear from you.