FAIRdata.ai: Making Open Data FAIR-er
What problem are we trying to solve?
Most major funders now require FAIR data management plans. Most generalist repositories (Zenodo, Figshare, Dryad, Mendeley Data) allow data published on them to be FAIR, but not all data published on them is FAIR. The gap between 'technically FAIR' and 'practically reusable' is large and I think there are ways to close it.
Visit any large repository and start browsing datasets at random. You'll find deposited data where the variable names are single letters, where the description field says 'data for paper', where there's no unit metadata, no controlled vocabulary alignment, no machine-readable schema.
FAIRdata.ai is trying to close that gap automatically. Given a dataset DOI, it assesses the current FAIR score across five independent frameworks, identifies what's missing, and then enriches the record - generating better descriptions, inferring variable semantics, aligning to standard ontologies, resolving licences to machine-readable SPDX URIs - without requiring anything from the original depositor. Every enrichment is tagged with full provenance so consumers know exactly what was added by whom (or what model), when, and with what confidence.
There's a second, less obvious problem FAIRdata.ai is working on. This is based on AllenAI’s work on Autodiscovery. I wrote about this here. Open datasets contain findings that have never been extracted. Correlations that nobody looked for. Distributions that are surprising given background knowledge. FAIRdata.ai surfaces these using Bayesian surprise scoring essentially asking 'what patterns in this dataset would be unexpected given domain priors?'. It feeds those signals upstream to OpenScience.ai as hypothesis seeds for formal investigation.
What tools have inspired or been used to build this platform?
FAIRdata.ai integrates a substantial number of open-source tools, open APIs, and community standards. Rather than building everything from scratch, the philosophy has been to compose existing infrastructure where it exists and build only where it doesn't. Below is a technical walkthrough of the stack, layer by layer.
Assessment: five FAIR frameworks, not one
Most platforms run a single FAIR checker. FAIRdata.ai runs five, then composites the results:
- F-UJI (FAIRsFAIR) - the most widely cited automated FAIR assessment tool. We self-host the F-UJI API and query it for 17 metrics across the four FAIR principles. F-UJI checks machine-readable metadata, persistent identifiers, licence declarations, data access protocols, and vocabulary usage.
- RDA FAIR Maturity Model - 41 indicators scored on a 0-5 maturity scale, from the Research Data Alliance FAIR Data Maturity Model Working Group. More granular than F-UJI, particularly strong on governance indicators.
- ARDC Self-Assessment - 12 practical checks from the Australian Research Data Commons. Pragmatic and well-calibrated for repository-deposited data.
- Metadata Game Changers - 58 documentation concepts covering both essential and supporting FAIR elements. Good at catching metadata completeness issues that other frameworks miss.
- Data Fitness for Use - an AI-assessed framework evaluating 14 reuse criteria from the perspective of a downstream consumer. This one uses an LLM to reason about whether a dataset is practically usable, not just technically compliant.
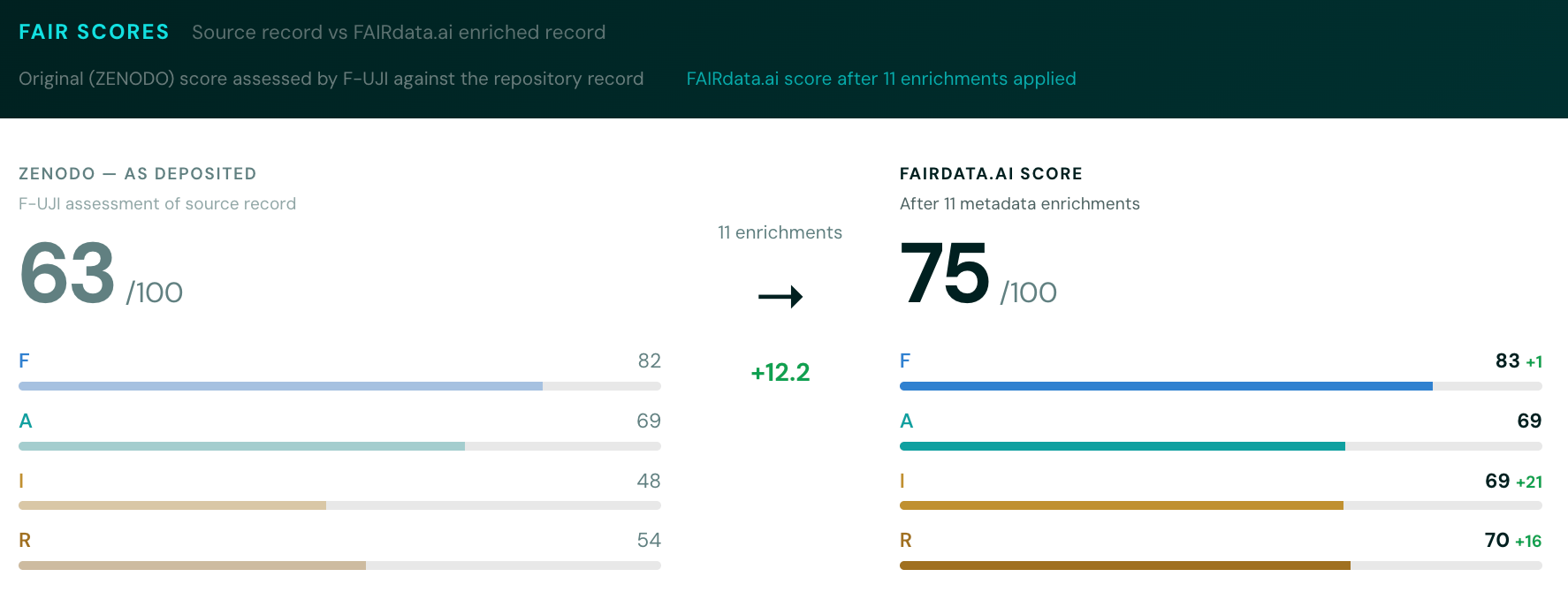
We run two passes: one on the source repository record as-is, one on the FAIRdata.ai-enriched version. The delta between these scores is arguably the most useful signal. It shows exactly how much better the record could be with automated enrichment.
AI-readiness scoring
Beyond FAIR, FAIRdata.ai scores datasets against two AI-readiness frameworks:
- GDS/DSIT 4-Pillar Framework (Jan 2026) - Technical Optimisation, Data & Metadata Quality, Organisation & Governance, Legal/Security/Ethics. This is the UK government's framework for assessing whether data is ready for AI/ML consumption.
- Bridge2AI AI-Readiness - 7 dimensions from the NIH Bridge2AI programme (Clark et al. 2024): FAIRness, Provenance, Characterisation, Pre-model Explainability, Ethics, Sustainability, Computability.
Metadata sources
Dataset metadata is normalised from multiple registries:
- DataCite Commons (api.datacite.org) - the primary metadata source. DataCite provides the canonical DOI metadata for most research data repositories.
- Crossref (api.crossref.org) - used for paper-to-dataset linking. When a dataset DOI has IsSupplementTo relationships or a linked publication DOI, we fetch the paper metadata to extract context.
Repository-specific APIs handle file listings and downloads:
- Figshare API (DOI prefix 10.6084)
- Zenodo API (DOI prefix 10.5281)
- Dryad API (DOI prefix 10.5061)
- Mendeley Data (DOI prefix 10.17632)
- Plus fallback patterns for Harvard Dataverse, OSF, GBIF, ICPSR, 4TU.ResearchData, ScienceDB, PhysioNet, and institutional Figshare instances.
Ontology and semantic enrichment
Raw metadata is enriched with controlled vocabulary annotations via:
- BioPortal (data.bioontology.org) we use three BioPortal endpoints: the Annotator API for extracting ontology terms from free text, the Search API for controlled vocabulary lookup, and the Recommender API for domain-aware ontology suggestions.
The ontologies we align to include:
- AGROVOC – agriculture, forestry, fisheries (FAO)
- MeSH – medical subject headings (NLM)
- ENVO – environmental ontology (biomes, ecosystems)
- Gene Ontology (GO) – molecular biology, gene function
- NCIT – NCI Thesaurus for biomedical terms
- CHEBI – chemical entities of biological interest
- EDAM – bioinformatics operations, data types, topics
- GAZ – geographic gazetteer
- SDGIO – Sustainable Development Goals Interface Ontology
Geographic enrichment uses Nominatim/OpenStreetMap for geocoding place names to coordinates.

AI enhancement pipeline
The metadata enrichment pipeline uses LLMs for tasks that require reasoning about semantics:
- Anthropic Claude (Sonnet) – the primary LLM for metadata enhancement. Generates descriptions, infers variable semantics, extracts paper context (methodology, limitations, population, ethics), and performs intelligent gap remediation. Every AI-generated enrichment is tagged with COMET provenance (agent, model, timestamp, confidence, derivation context).
The enhancement is orchestrated through a chain of specialised agents:
- MetadataHarvester – extracts structured metadata from unstructured sources
- PaperLinker – finds and links related publications via Crossref and OpenAlex
- PaperContextExtractor – extracts methodology, limitations, and domain context from linked papers
- SubjectEnricher – enhances subject keywords with ontology-aligned terms
- OntologyAnnotator – maps free-text terms to BioPortal ontology concepts
- GeoEntityExtractor – identifies and geocodes geographic references
- TemporalExtractor – identifies temporal coverage and resolution
- FormatDetector – identifies file formats and suggests MIME types
- LicenseResolver – maps licence text to machine-readable SPDX URIs
- SemanticResourceGenerator -- generates schema.org JSON-LD markup
- IntelligentRemediation – fills remaining metadata gaps
- ProvenanceBuilder – assembles COMET provenance records for all enrichments
- QualityStatementGenerator – produces human-readable quality assessments
Data profiling
For datasets containing tabular files (CSV, TSV, Excel, Parquet), FAIRdata.ai performs column-level statistical profiling using pandas, NumPy, and SciPy:
- Column data types, cardinality, null rates
- Distributions (mean, median, std, skewness, quartiles)
- Outlier detection and quality flags
- Top values for categorical columns
These profiles power both the discovery engine (identifying interesting statistical patterns) and the cross-dataset matching system (finding datasets with shared column structure).
Discovery engine

The discovery layer is where FAIRdata.ai goes beyond assessment into knowledge generation:
- AutoDiscovery MCTS sidecar -- a Monte Carlo Tree Search engine that explores the hypothesis space of a dataset's statistical properties. It uses OpenAI GPT-4o for reasoning about which analyses to run, then executes them on the actual data. Each finding is scored with Bayesian surprise (prior-to-posterior shift) to measure genuine novelty rather than mere statistical significance. Analyses include correlation detection, distribution characterisation, bimodality testing, group differences (ANOVA), outlier classification, and temporal trend detection.
- Cross-dataset discovery -- a system that finds joinable dataset pairs by matching column structure across the profiled dataset corpus. When two datasets share meaningful columns (filtering out garbage like Unnamed: 0 or generic fields like id), they can be merged and submitted to the MCTS engine for cross-study hypothesis generation.
High-confidence findings (Bayesian surprise >= 0.8, p < 0.01) are automatically promoted and pushed to OpenScience.ai as research seeds, where they're processed into formal hypothesis papers.
Packaging and export formats
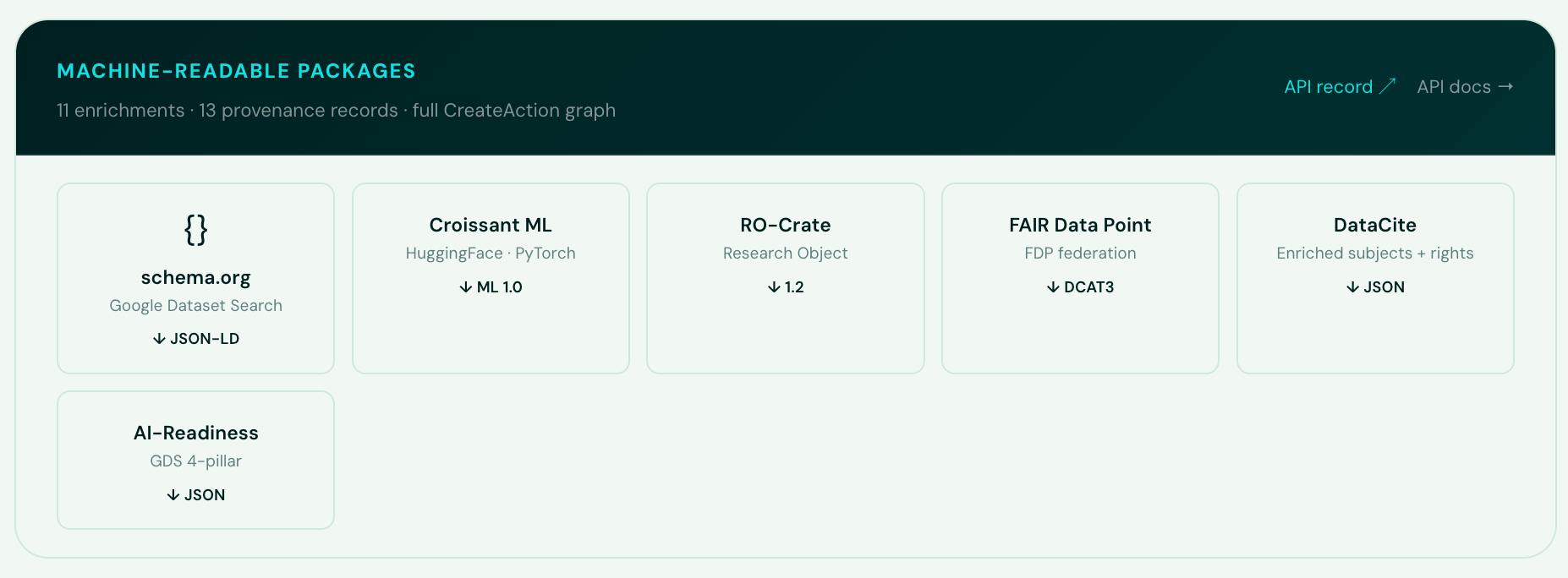
Enriched records are packaged into six machine-readable formats using FAIRSCAPE:
- schema.org/Dataset – JSON-LD for Google Dataset Search discoverability
- Croissant ML 1.0 – the MLCommons standard for ML-ready datasets (compatible with PyTorch, TensorFlow, JAX, HuggingFace, Kaggle)
- RO-Crate 1.2 – Research Object packaging with full provenance
- FAIR Data Point (DCAT3) – Turtle RDF for FAIR data exchange
- Enriched DataCite JSON – structured subjects with ontology URIs, SPDX rights
- AI-Readiness JSON – GDS/DSIT + Bridge2AI scoring
Agent integration
FAIRdata.ai exposes its capabilities to AI agents via:
- Model Context Protocol (MCP) –JSON-RPC 2.0 endpoint at /mcp for Claude, ChatGPT, and custom agent integration
- OpenAI Plugin Manifest – /.well-known/ai-plugin.json for ChatGPT plugin discovery
- llms.txt – standardised LLM-readable documentation at /llms.txt
Infrastructure
- Backend: FastAPI (Python 3.11, async) with Uvicorn, deployed on Railway
- Frontend: React with Vite, Recharts for data visualisation, deployed on Netlify
- Database: Supabase (managed PostgreSQL)
- Analytics: Plausible (privacy-friendly, cookie-free)
- Provenance: COMET model -- every AI enrichment carries agent ID, model version, timestamp, confidence score, and derivation context
What we can look to do next
The most obvious gap is coverage. FAIRdata.ai currently works well with Figshare, Zenodo, Dryad, Harvard Dataverse, Mendeley Data, OSF, and Vivli, but the tail of domain repositories is long and each has its own metadata quirks.
There are several technical directions I'm exploring:
The two-score nudge. The gap between a dataset's original FAIR score and its enriched score is a meaningful signal. Showing depositors that their record went from 62% to 84% with automated enrichment -- and exactly which fields were improved -- could be a practical nudge toward better data curation practice at the point of deposit.Stronger discovery-to-hypothesis pipeline. Right now the Bayesian surprise scoring identifies datasets with interesting distributional properties, but the handoff to OpenScience.ai is still coarse. Tightening this so FAIRdata.ai passes not just 'this dataset is interesting' but 'here is a specific, testable claim with statistical evidence and domain context' would make the whole pipeline considerably more powerful.
Federated assessment. Rather than pulling datasets to our infrastructure, enabling repositories to run FAIRdata.ai assessment in-place (as a service or plugin) would scale coverage without scaling compute. The F-UJI framework already supports this model; extending it to the full five-framework composite would be the contribution.
If you're working on any of these problems and want to compare notes, or if you spot something obviously wrong with the approach, I'd be glad to hear from you.