What parts of the academic knowledge creation & dissemination pipeline can we automate with AI?
As a tech founder who can, at best, describe himself as a hacky coder, I've spent most of my career having to either hire engineers or do without when an idea was too technically demanding. That's changed quite a lot recently. Tools like Claude Code mean I can now actually build the things I want to test, which makes experimenting in this space genuinely good fun in a way it wouldn't have been two or three years ago. So with that said - what parts of the academic knowledge creation and dissemination pipeline can we automate thanks to AI? Below, I describe 4 platforms that try to individually answer parts of this question, and when combined, attempt to answer a little bit more of it.
As a disclaimer, everything discussed here is experimental. The platforms described are not robust scholarly infrastructure. They should not be cited, relied upon, or treated as production systems. They're a hobby project built on open APIs, open data, and open infrastructure designed to test what's technically possible.
The problems worth solving
We've built good infrastructure for sharing research outputs, yet most of the value locked in those outputs goes unrealised. Meanwhile, the research community is under pressure in almost every dimension: time, funding, reviewer bandwidth, editorial capacity. A disproportionate amount of effort goes on tasks that are fairly mechanical - formatting metadata, screening submissions for integrity, summarising prior literature, running standard statistical checks.
New academic knowledge creation and basic research has never been more in demand. AI will never replace this. But there is a subset of academic knowledge gap filling that new tools seem capable of filling. Can they? What if some of those tasks could be automated? I am not attempting here answer the question of “What should be automated?”
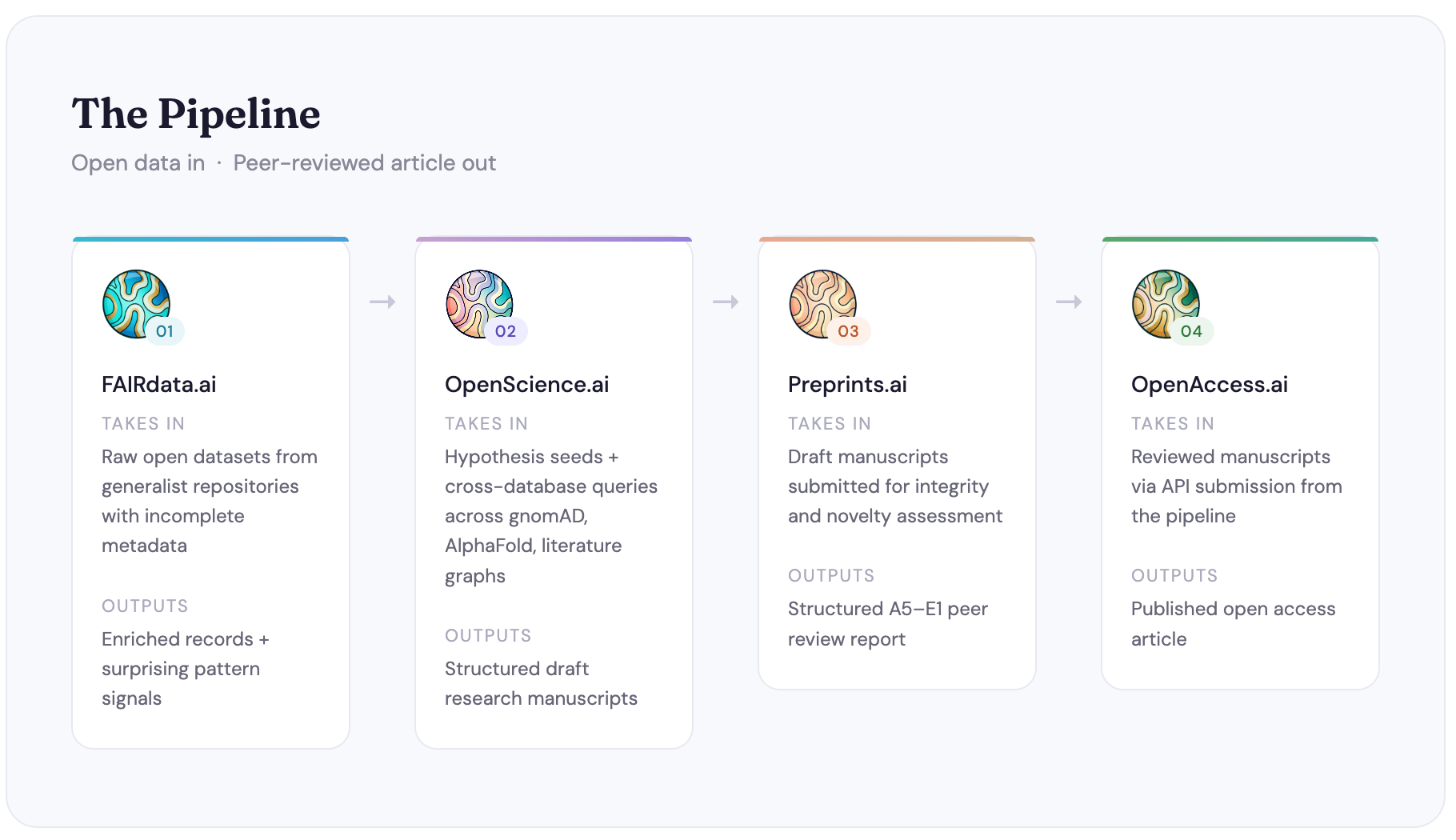
The four platforms, briefly

FAIRdata.ai starts at the data layer. A large proportion of datasets deposited on generalist repositories are technically FAIR but practically unusable. The metadata is incomplete, the variable labels are opaque, the context is missing. FAIRdata.ai tries to fix that automatically, enriching metadata to improve FAIR scores. It also tries to make use of surprisal scoring and automated discovery techniques to surface findings within datasets that may never have been noticed by the original depositors. Those signals feed into OpenScience.ai.You can read more details about the technical capabilities and dependencies of FAIRdata.ai here.
OpenScience.ai is the hypothesis engine. It queries open scientific databases. eg. gnomAD, Ensembl, AlphaFold, the literature graph, and any hypotheses that FAIRdata.ai sends it and tries to identify patterns that haven't yet been written up as research articles. It then generates hypotheses, assembles supporting evidence, and drafts those hypotheses into structured manuscripts. It's not trying to replace the experimental scientist; it's trying to explore the space of already-existing open data for combinations that nobody has looked at yet.
You can read more details about the technical capabilities and dependencies of OpenScience.ai here.
Preprints.ai asks a fairly direct question - How much of peer review can we automate? It runs a multi-agent review pipeline assessing both research integrity, methodology, statistical validity, reproducibility, and novelty. The output is a structured assessment graded against a rubric, with deliberative rounds between agents before a consensus is reached.
You can read more details about the technical capabilities and dependencies of Preprints.ai here.
OpenAccess.ai closes the loop. If we're generating articles via OpenScience.ai and reviewing them via Preprints.ai, we need somewhere to publish them. OpenAccess.ai is that place - an open access journal built on the assumption that the editorial process can run largely on agents, and that the unit cost of academic publishing can be meaningfully reduced as a result.
You can read more details about the technical capabilities and dependencies of OpenAccess.ai here.
All four platforms are built exclusively on open data and open APIs. If you can build a plausible end-to-end research automation pipeline using only what's publicly available, that says something useful about the current state of open science infrastructure. The constraint on scientific productivity isn't data availability; it's the capacity for machines to process and synthesise it.
I personally have access to considerably richer data through my work at Digital Science (citation graphs, full-text repositories, institutional metadata). I haven't used any of it here. What you're seeing is what's possible with no proprietary advantage, which I think makes the findings more generalisable.
Humans in the loop
Framing this as "minimal human intervention" is accurate for certain parts of the pipeline and misleading for others.
There are stages where the automation is fairly robust. Metadata enrichment, FAIR scoring, integrity screening, novelty checks against the existing literature. These are tasks where the output is verifiable and the failure modes are containable. They're also, not coincidentally, the tasks that consume a lot of researcher time without requiring much researcher judgement.
But there are other stages where a human needs to be in the loop, and probably will for a while. Editorial decisions - whether a piece of work is actually worth publishing, whether the framing is intellectually honest, whether the conclusions follow from the evidence. These are not tasks I'd be comfortable delegating to agents at this point. The current OpenAccess.ai setup allows human editors to review agent assessments before making final decisions, and I think that hybrid model is the right one for this experiment. It also leans very heavily on eLife’s Publish, Review, Curate model, which is a model I see gaining more traction and uptake in the coming years.
Similarly, the hypothesis generation in OpenScience.ai needs a human sanity check before anything gets treated as a serious research direction. The system can identify patterns in open data, but it can't yet reliably distinguish a genuinely interesting finding from a statistical artefact or a confound that any domain expert would spot immediately. I've had to be quite disciplined about this after discovering early on that the pipeline would confidently produce fabricated statistics if you didn't anchor every claim to verifiable data from a canonical source.
What's next
Each platform has a reasonably long list of things it still can't do well. OpenScience.ai's anti-fabrication infrastructure is solid now but the discovery logic is still fairly shallow. Preprints.ai's local inference falls over on structured output for complex multi-agent workflows. FAIRdata.ai's scoring is imprecise for accession-based repositories. OpenAccess.ai's cost economics need testing at meaningful volume before any of the unit cost claims are worth much.
I'm writing this up now in order to experiment in the open. The idea of chaining these capabilities together is the more interesting thing to put into the world, even if the implementations are still rough.
If you're working on any of these problems and want to compare notes, or if you spot something obviously wrong with the approach, I'd be glad to hear from you.